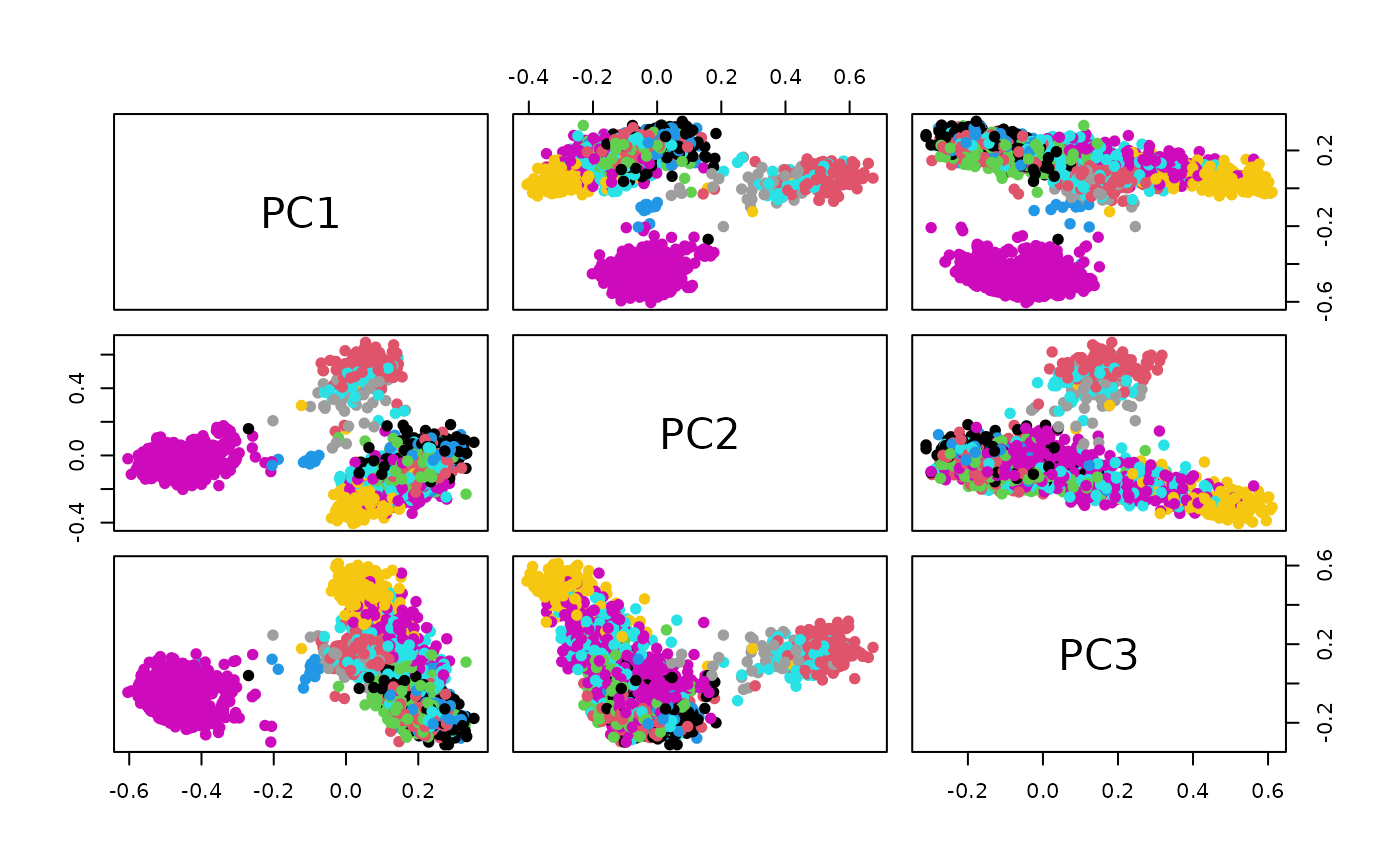
update a SingleCellExperiment with PCA (via irlba) reductions of embeddings produced with PyTorch-BigGraph
Source:R/dimreds.R
pca_CG.Rdupdate a SingleCellExperiment with PCA (via irlba) reductions of embeddings produced with PyTorch-BigGraph
pca_CG(trout, sce, ncomp = 3, ...)Arguments
- trout
output of `BiocPBG::train_eval`
- sce
SingleCellExperiment instance derived from or consistent with the one from which `trout` was produced
- ncomp
numeric(1) passed as `n` to `irlba::prcomp_irlba`
- ...
named arguments other than `n` to be passed to irlba
Value
The `sce` is updated to include a `reducedDim` component named `PCA` based on the cell embedding, and the rowData component is updated to include the PCA applied to gene embedding