uberonpeek -- a look at UBERON ontology, etc., with ontoProc2
Vincent J. Carey, stvjc at channing.harvard.edu
March 25, 2026
Source:vignettes/uberonpeek.Rmd
uberonpeek.RmdIntroduction
The ontoProc2 package is designed to give convenient access to the ontologies that are transformed to “semantic SQL” in the INCAtools project.
We’ll start by retrieving the current UBERON ontology and examining some tables and “statements”.
library(ontoProc2)
library(DBI)
library(dplyr)
ubss = semsql_connect(ontology="uberon")
report(ubss)##
## ============================================================
## SemsqlConn Object
## ============================================================
##
## Connection Details:
## ----------------------------------------
## Database path: /Users/vincentcarey/Library/Caches/org.R-project.R/R/BiocFileCache/40e22de1c3cc_uberon.db
## Ontology prefix: UBERON
## Status: ✓ Connected
##
## Database Statistics:
## ----------------------------------------
## Labeled terms: 28,420
## Direct edges: 80,252
## Entailed edges: 5,312,269
## Definitions: 22,660
##
## Terms by Prefix (top 5):
## ----------------------------------------
## UBERON: 15,770
## GO: 7,428
## CL: 1,473
## _: 1,260
## CHEBI: 915
##
## Key Tables Available:
## ----------------------------------------
## ✓ rdfs_label_statement
## ✓ has_text_definition_statement
## ✓ edge
## ✓ entailed_edge
## ✓ rdfs_subclass_of_statement
## ✓ owl_some_values_from
## ✓ has_oio_synonym_statement
##
## ============================================================
## Use methods like search_labels(), get_ancestors(), etc.
## Run ?SemsqlConn for documentation.
## ============================================================
ubcon = ubss@con
head(dbListTables(ubcon))## [1] "all_problems" "annotation_property_node"
## [3] "anonymous_class_expression" "anonymous_expression"
## [5] "anonymous_individual_expression" "anonymous_property_expression"
tbl(ubcon, "statements")## # Source: table<`statements`> [?? x 8]
## # Database: sqlite 3.51.2 [/Users/vincentcarey/Library/Caches/org.R-project.R/R/BiocFileCache/40e22de1c3cc_uberon.db]
## stanza subject predicate object value datatype language graph
## <chr> <chr> <chr> <chr> <chr> <chr> <chr> <chr>
## 1 obo:uberon.owl obo:uberon.owl foaf:home… NA http… xsd:any… NA NA
## 2 obo:uberon.owl obo:uberon.owl rdfs:comm… NA Aure… NA NA NA
## 3 obo:uberon.owl obo:uberon.owl oio:treat… NA ZFS … NA NA NA
## 4 obo:uberon.owl obo:uberon.owl oio:treat… NA ZFA … NA NA NA
## 5 obo:uberon.owl obo:uberon.owl oio:treat… NA XAO … NA NA NA
## 6 obo:uberon.owl obo:uberon.owl oio:treat… NA WBls… NA NA NA
## 7 obo:uberon.owl obo:uberon.owl oio:treat… NA WBbt… NA NA NA
## 8 obo:uberon.owl obo:uberon.owl oio:treat… NA TGMA… NA NA NA
## 9 obo:uberon.owl obo:uberon.owl oio:treat… NA TAO … NA NA NA
## 10 obo:uberon.owl obo:uberon.owl oio:treat… NA TADS… NA NA NA
## # ℹ more rowsParent-child relations
CRAN’s ontologyIndex package provides a familiar representation that simplifies visualization.
uboi = semsql_to_oi(ubcon)## Warning in ontologyIndex::ontology_index(name = nn, parents = pl): Some parent
## terms not found: BFO:0000001, CARO:0000000, CHEBI:24431 (5 more)
uboi## Ontology with 25523 terms
##
## Properties:
## id: character
## name: list
## parents: list
## children: list
## ancestors: list
## obsolete: logical
## Roots:
## CHEBI:24432 - biological role
## CHEBI:51086 - chemical role
## CHEBI:33232 - application
## CHEBI:23367 - molecular entity
## CHEBI:24433 - group
## BFO:0000002 - continuant
## CHEBI:33250 - atom
## BFO:0000003 - occurrent
## CARO:0000007 - immaterial anatomical entity
## CHEBI:36340 - fermion
## ... 9 more
uboi$name[10364:10370]## $`PATO:0000070`
## [1] "amount"
##
## $`PATO:0000136`
## [1] "closure"
##
## $`PATO:0000141`
## [1] "structure"
##
## $`PATO:0000150`
## [1] "texture"
##
## $`PATO:0000169`
## [1] "viability"
##
## $`PATO:0000261`
## [1] "maturity"
##
## $`PATO:0000322`
## [1] "red"A sense of the variety of ontological cross-references present can be given by tabling the tag prefixes.
## prefs
## BFO BSPO CARO CHEBI CL GO IAO NBO
## 14 12 5 912 1470 7427 5 37
## NCBITaxon PATO PR RO UBERON
## 473 159 333 1 14675By using the ancestors component we can obtain a view of is-a relations (presumably developed from rdfs:subClassOf predicates). We’ve chosen as terminal tags the tags for heart, kidney, and cortex of kidney.
onto_plot2(uboi,
unlist(uboi$ancestors[c("UBERON:0002189",
"UBERON:0002113", "UBERON:0000948")]))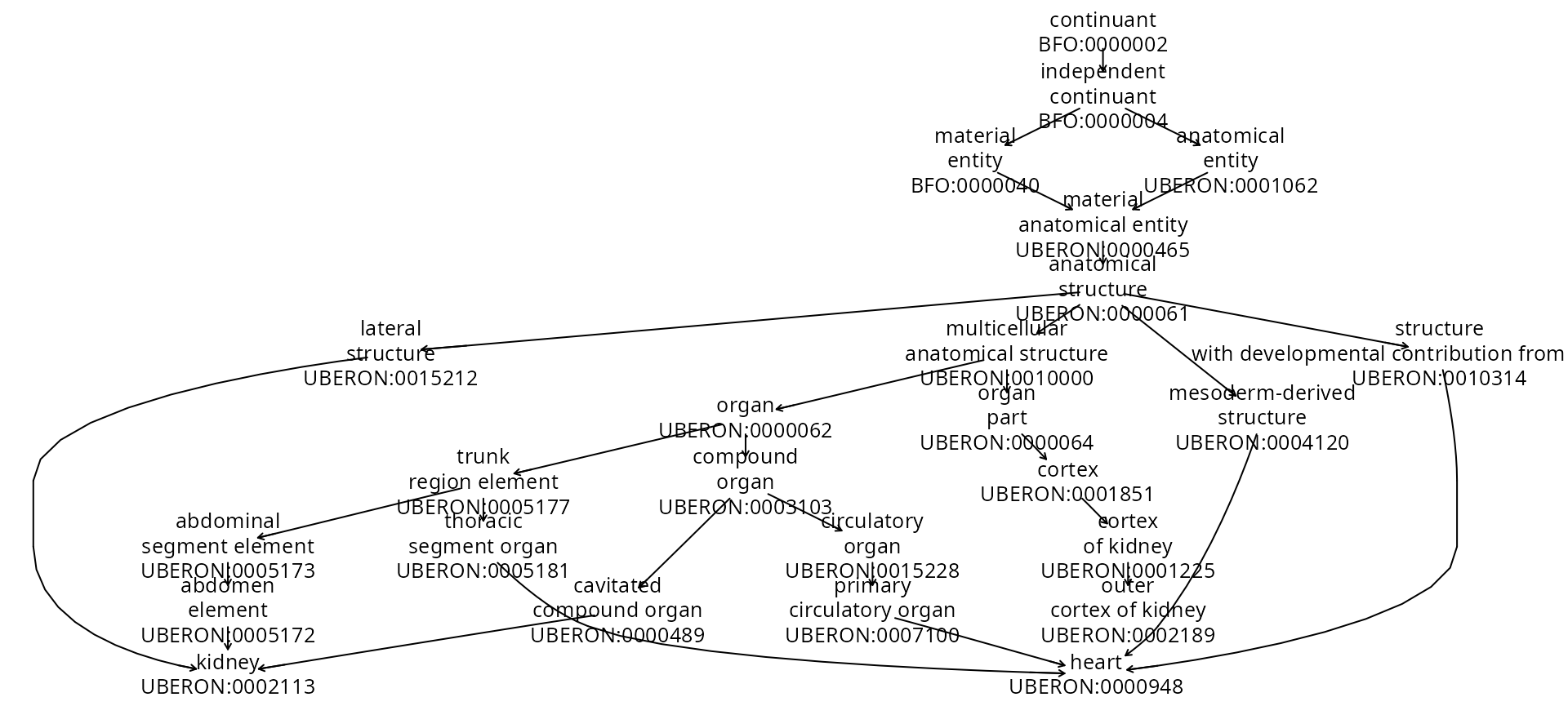
EFO and NCI thesaurus
On cursory inspection, the EFO ontology has considerable information about anatomic locations of diseases.
We’ll use the entailed edges table in EFO to find all statements that have ‘heart’ (UBERON:0000948) as object.
eforef = semsql_connect(ontology="efo") # 240 MB## Connected to SemanticSQL database: /Users/vincentcarey/Library/Caches/org.R-project.R/R/BiocFileCache/41b765188508_efo.db## Primary ontology prefix: EFO
#nciref = semsql_connect("ncit") # > 500MB, block
htab = tbl(eforef@con, "entailed_edge") |>
filter(object == "UBERON:0000948") |> as.data.frame()
head(htab)## subject predicate object
## 1 UBERON:0000948 rdfs:subClassOf UBERON:0000948
## 2 OBA:VT0000266 RO:0002502 UBERON:0000948
## 3 EFO:0003896 RO:0002502 UBERON:0000948
## 4 OBA:2050102 RO:0002502 UBERON:0000948
## 5 OBA:0006021 RO:0002502 UBERON:0000948
## 6 OBA:VT0004063 RO:0002502 UBERON:0000948It is tedious to see these formal tags. We have assembled a simple character vector map that covers many tags.
## IAO:0000112 IAO:0000114 IAO:0000115
## "example of usage" "has curation status" "definition"
## IAO:0000116 IAO:0000117 IAO:0000232
## "editor note" "term editor" "curator note"What are the predicates of the heart table above?
ncit_map[unique(htab$predicate)]## <NA> RO:0002502
## NA "depends on"
## IAO:0000136 <NA>
## "is_about" NA
## BFO:0000066 RO:0001025
## "occurs in" "located_in"
## RO:0002131 RO:0002314
## "overlaps" "inheres in part of"
## EFO:0000784 BFO:0000050
## "has_disease_location" "part_of"
## RO:0000052 RO:0004027
## "inheres_in" "disease has inflammation site"To enumerate and decode the terms with disease location (EFO:0000784) in heart, we have
Session information
## R version 4.5.2 (2025-10-31)
## Platform: aarch64-apple-darwin20
## Running under: macOS Sequoia 15.7.4
##
## Matrix products: default
## BLAS: /System/Library/Frameworks/Accelerate.framework/Versions/A/Frameworks/vecLib.framework/Versions/A/libBLAS.dylib
## LAPACK: /Library/Frameworks/R.framework/Versions/4.5-arm64/Resources/lib/libRlapack.dylib; LAPACK version 3.12.1
##
## locale:
## [1] en_US.UTF-8/en_US.UTF-8/en_US.UTF-8/C/en_US.UTF-8/en_US.UTF-8
##
## time zone: America/New_York
## tzcode source: internal
##
## attached base packages:
## [1] stats graphics grDevices utils datasets methods base
##
## other attached packages:
## [1] DT_0.34.0 dplyr_1.2.0 DBI_1.3.0 ontoProc2_0.99.8
## [5] BiocStyle_2.38.0
##
## loaded via a namespace (and not attached):
## [1] utf8_1.2.6 rappdirs_0.3.4 sass_0.4.10
## [4] generics_0.1.4 RSQLite_2.4.6 digest_0.6.39
## [7] magrittr_2.0.4 evaluate_1.0.5 grid_4.5.2
## [10] bookdown_0.46 fastmap_1.2.0 blob_1.3.0
## [13] R.oo_1.27.1 jsonlite_2.0.0 ontologyIndex_2.12
## [16] R.utils_2.13.0 ontologyPlot_1.7 graph_1.88.1
## [19] BiocManager_1.30.27 purrr_1.2.1 crosstalk_1.2.2
## [22] Rgraphviz_2.54.0 httr2_1.2.2 textshaping_1.0.4
## [25] jquerylib_0.1.4 paintmap_1.0 cli_3.6.5
## [28] rlang_1.1.7 dbplyr_2.5.2 R.methodsS3_1.8.2
## [31] bit64_4.6.0-1 withr_3.0.2 cachem_1.1.0
## [34] yaml_2.3.12 otel_0.2.0 tools_4.5.2
## [37] memoise_2.0.1 filelock_1.0.3 BiocGenerics_0.56.0
## [40] curl_7.0.0 vctrs_0.7.1 R6_2.6.1
## [43] stats4_4.5.2 BiocFileCache_3.0.0 lifecycle_1.0.5
## [46] fs_1.6.6 htmlwidgets_1.6.4 bit_4.6.0
## [49] ragg_1.5.0 pkgconfig_2.0.3 desc_1.4.3
## [52] pkgdown_2.2.0 pillar_1.11.1 bslib_0.10.0
## [55] glue_1.8.0 systemfonts_1.3.1 xfun_0.56
## [58] tibble_3.3.1 tidyselect_1.2.1 knitr_1.51
## [61] htmltools_0.5.9 rmarkdown_2.30 compiler_4.5.2
## [64] S7_0.2.1